back to home
Research
My research has focused on adapting deep learning models to the complexity of biological data through novel architectures, regularizations, and training setups. I'm especially interested in representation learning and reasoning models.
For a full publication list, see my Google Scholar profile.
Selected Projects
Newest work appears first.
|
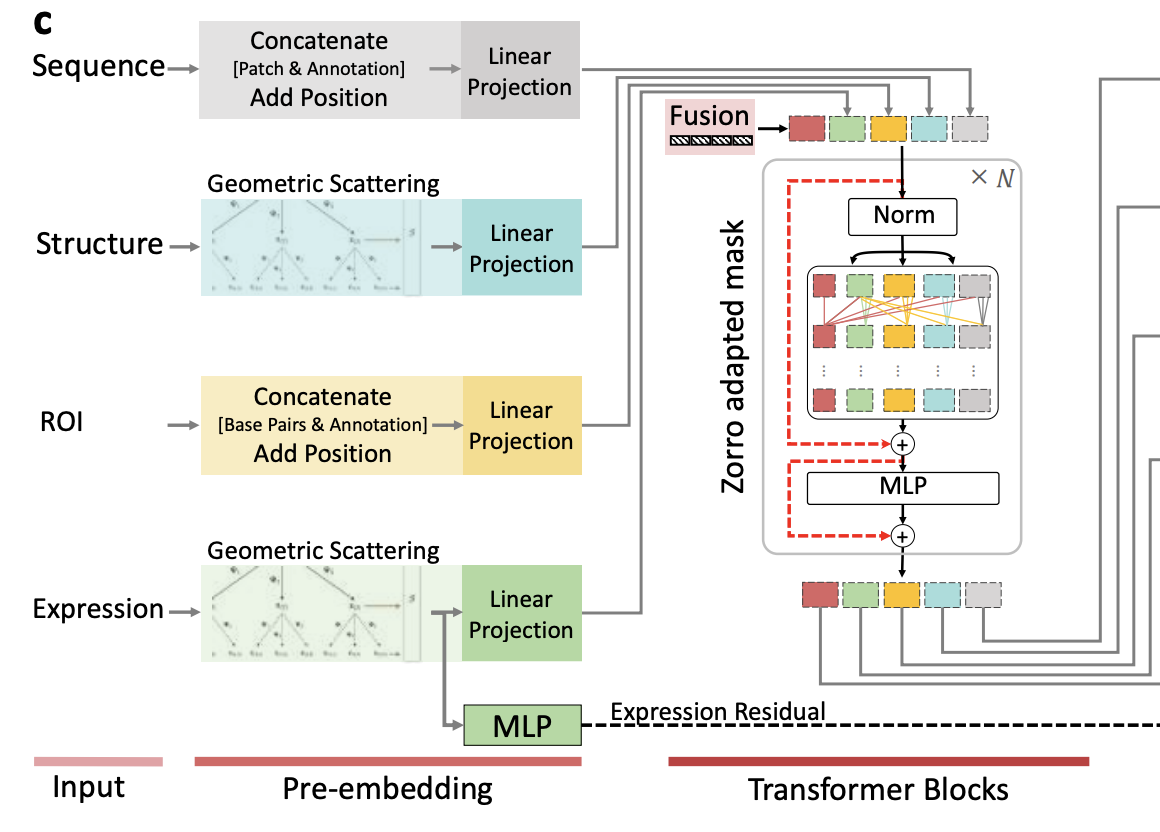
2025 · CellSpliceNet: interpretable multimodal modeling of alternative splicing across neurons
CellSpliceNet develops interpretable multimodal models of alternative splicing across C. elegans neurons, integrating complementary biological signals to characterize neuron-specific splicing programs. bioRxiv preprint | NeurIPS workshop paper |
|
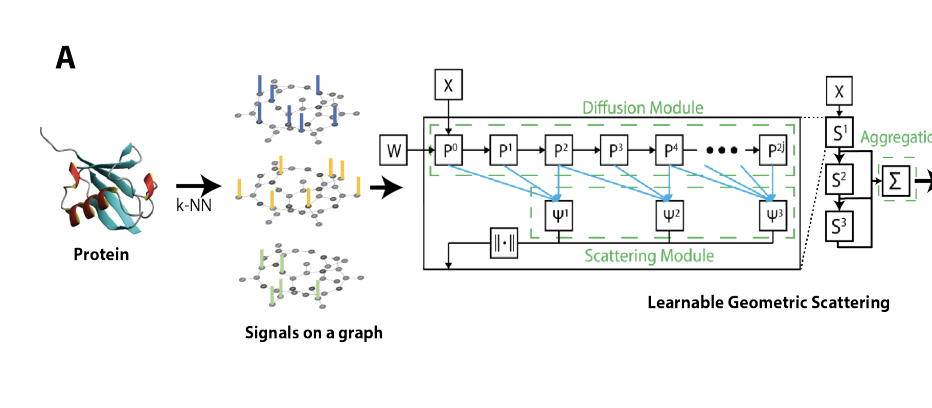
2024 · ProtSCAPE: mapping the landscape of protein conformations in molecular dynamics
ProtSCAPE develops representation-learning methods for organizing protein conformations from molecular dynamics simulations, making structural ensembles easier to map, compare, and interpret. arXiv preprint | conference presentation | Presented at SMT Conference, 2025 |
|
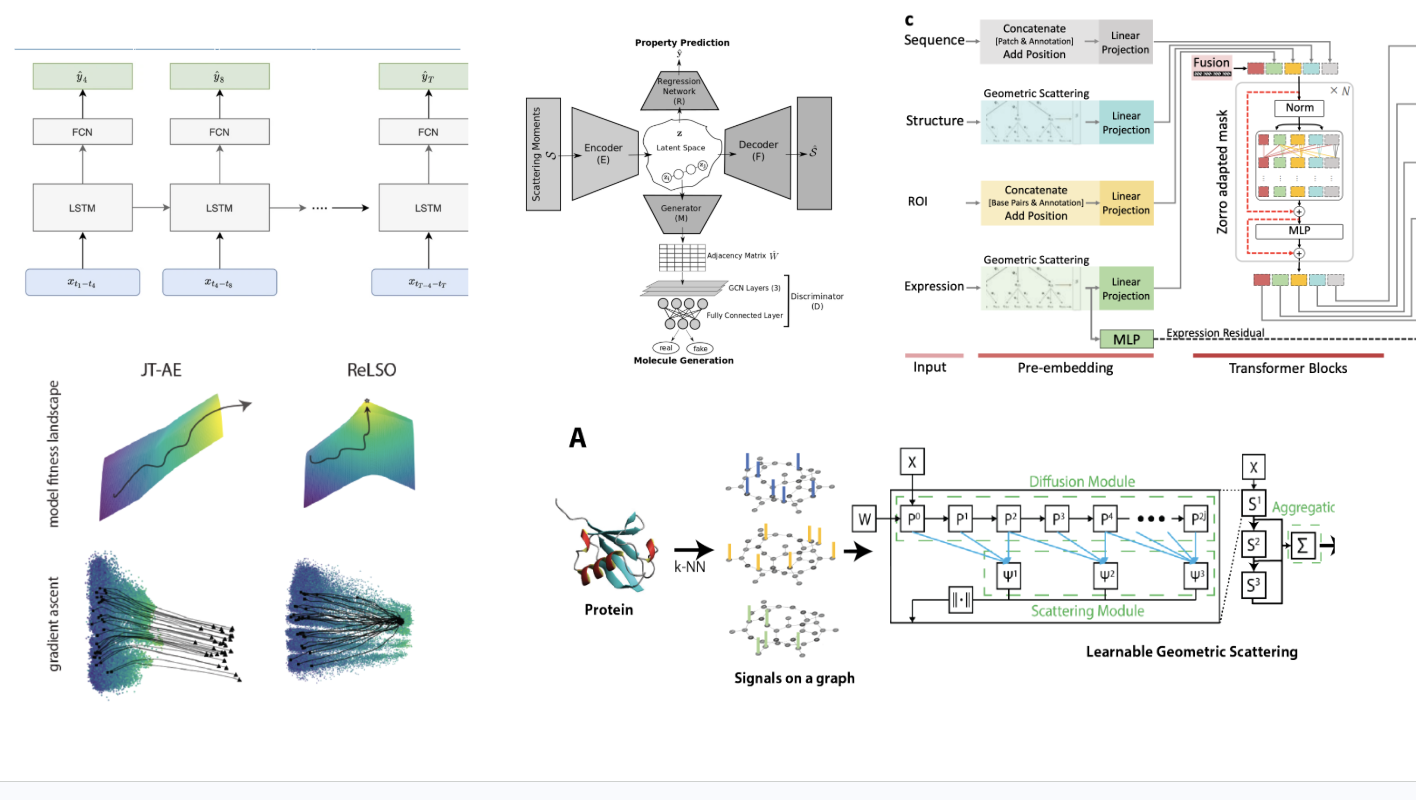
2023 · Representation learning and generative modeling for biomolecular sequences and structures
My dissertation unifies work on representation learning, generative modeling, and biomolecular optimization across proteins, RNA, and molecular graphs, with an emphasis on learned representations that support both scientific analysis and design. dissertation |
|
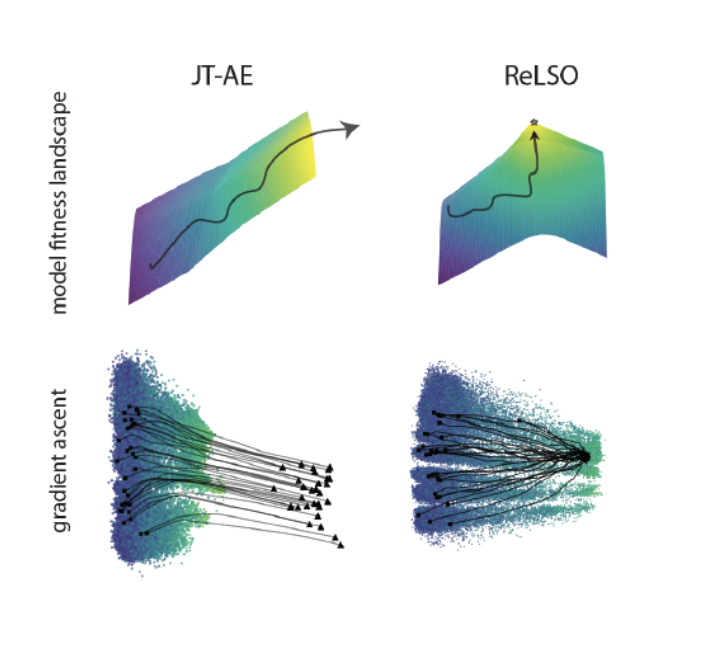
2022 · Transformer-based protein generation with regularized latent-space optimization
This work develops transformer-based protein generative models with regularized latent spaces, enabling gradient-guided search in learned sequence representations for more controllable protein design and optimization. Nature Machine Intelligence paper | ReLSO preprint | Presented at ISMB/ECCB 2021, the Learning Meaningful Representations of Life Workshop at ICLR 202 |
|
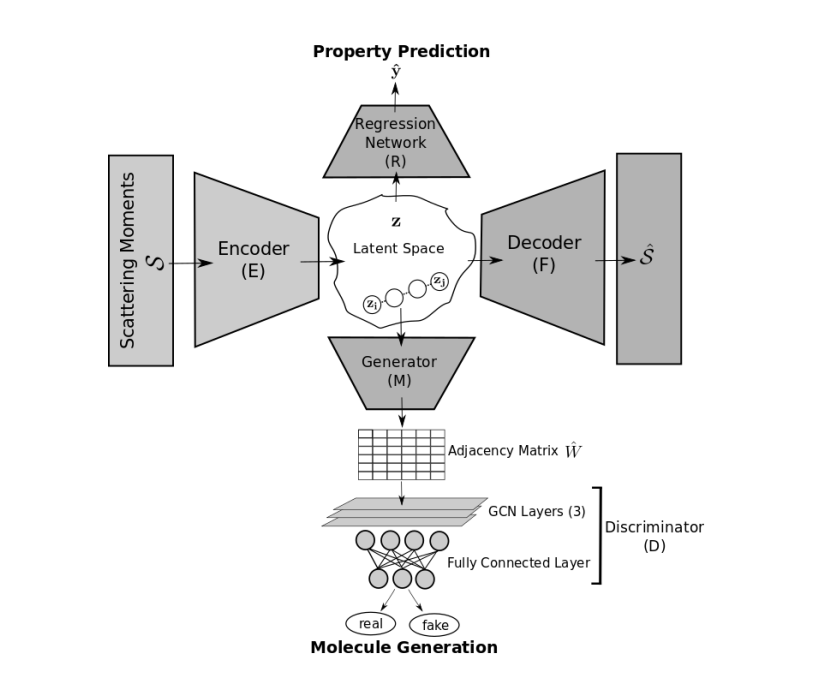
2022 · Molecular graph generation via geometric scattering
This project combines geometric scattering with generative modeling for molecules, testing whether graph-based signal-processing representations can capture chemically meaningful structure for molecular generation. conference paper | Presented at IEEE MLSP, 2022 |
|
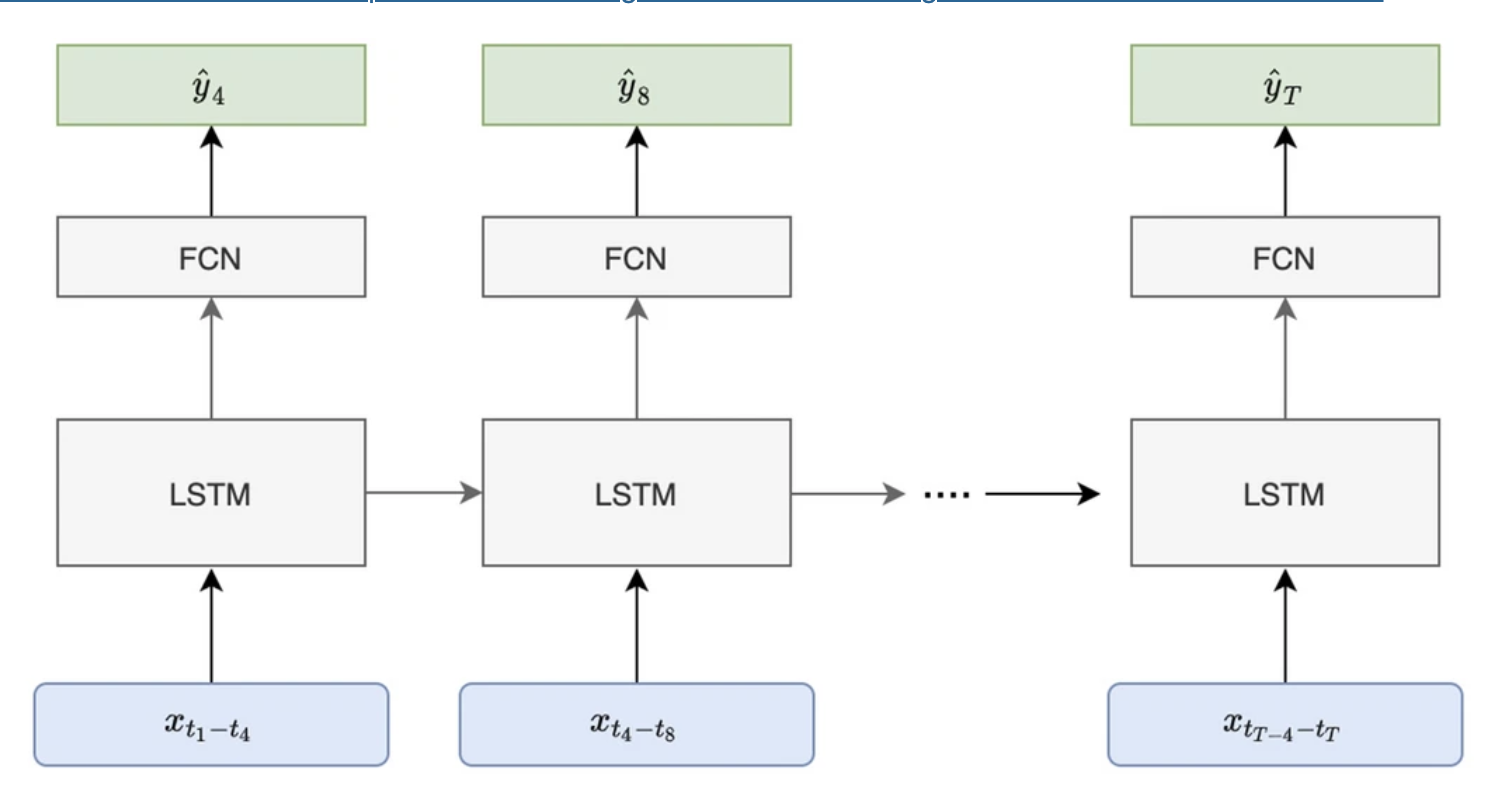
2021 · Predicting transfusion need in acute GI bleeding
This clinical machine learning project develops neural-network models to predict whether ICU patients with acute gastrointestinal bleeding will require red blood cell transfusion, supporting earlier risk stratification in high-acuity care settings. Scientific Reports paper | conference abstract |
|
|
2020 · Uncovering the folding landscape of RNA secondary structure using deep graph embeddings
This project uses deep graph embeddings to recover structure in RNA folding landscapes, connecting RNA secondary-structure analysis with representation learning over graph-based molecular objects. IEEE Big Data paper | arXiv preprint | Presented at the GRLB Workshop at ICML and at IEEE Big Data, 2020 |
|
|
2017 · Vitamin K epoxide reductase regulation of androgen receptor activity
This earlier work examined how vitamin K epoxide reductase regulates androgen receptor activity, contributing to mechanistic understanding relevant to cancer biology and translational research. journal paper |